Designing information-dense promoters by packing overlapping binding sites
De novo promoter sequence design via nucleotide string packing and integer linear programming.
- Cis-regulatory DNA
- Sequence design algorithms
- Integer linear programming
- promoter design
Promoters control when and how genes are transcribed. In simple constitutive expression, RNA polymerase activity is mostly set by basal promoter strength. In regulated expression, multiple transcription factors shape initiation through their binding, activity, and effective occupancy, often in a condition-dependent way.
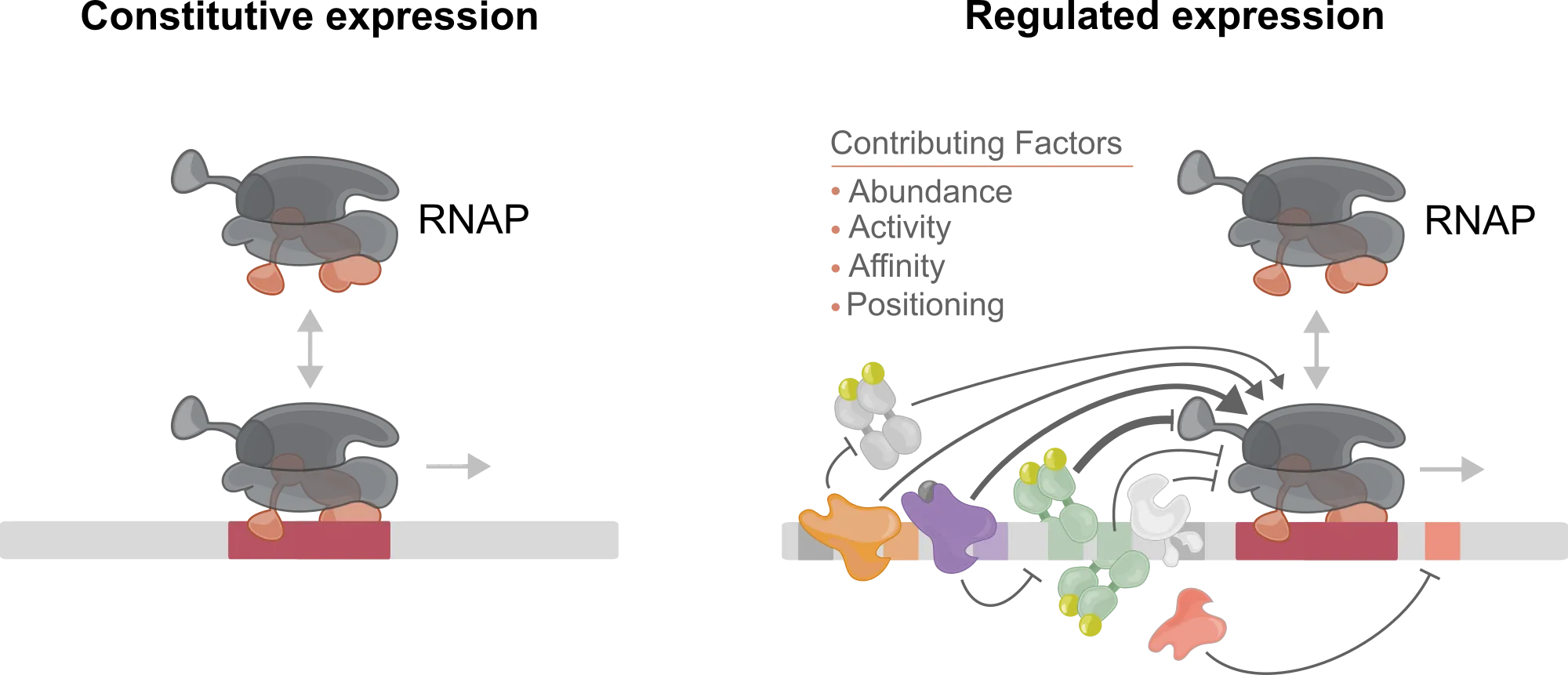
How do you design that kind of regulatory complexity from scratch, especially when binding sites can overlap and compete within a short sequence?
I helped develop dense-arrays, an
in silico approach that frames this as a nucleotide string-packing problem.
The method packs many DNA–protein binding sites into compact, contiguous arrays
by reducing the search to an optimization problem solved with integer linear
programming, which makes it practical to generate either diverse libraries or
focused constructs without hand-tuning each architecture.
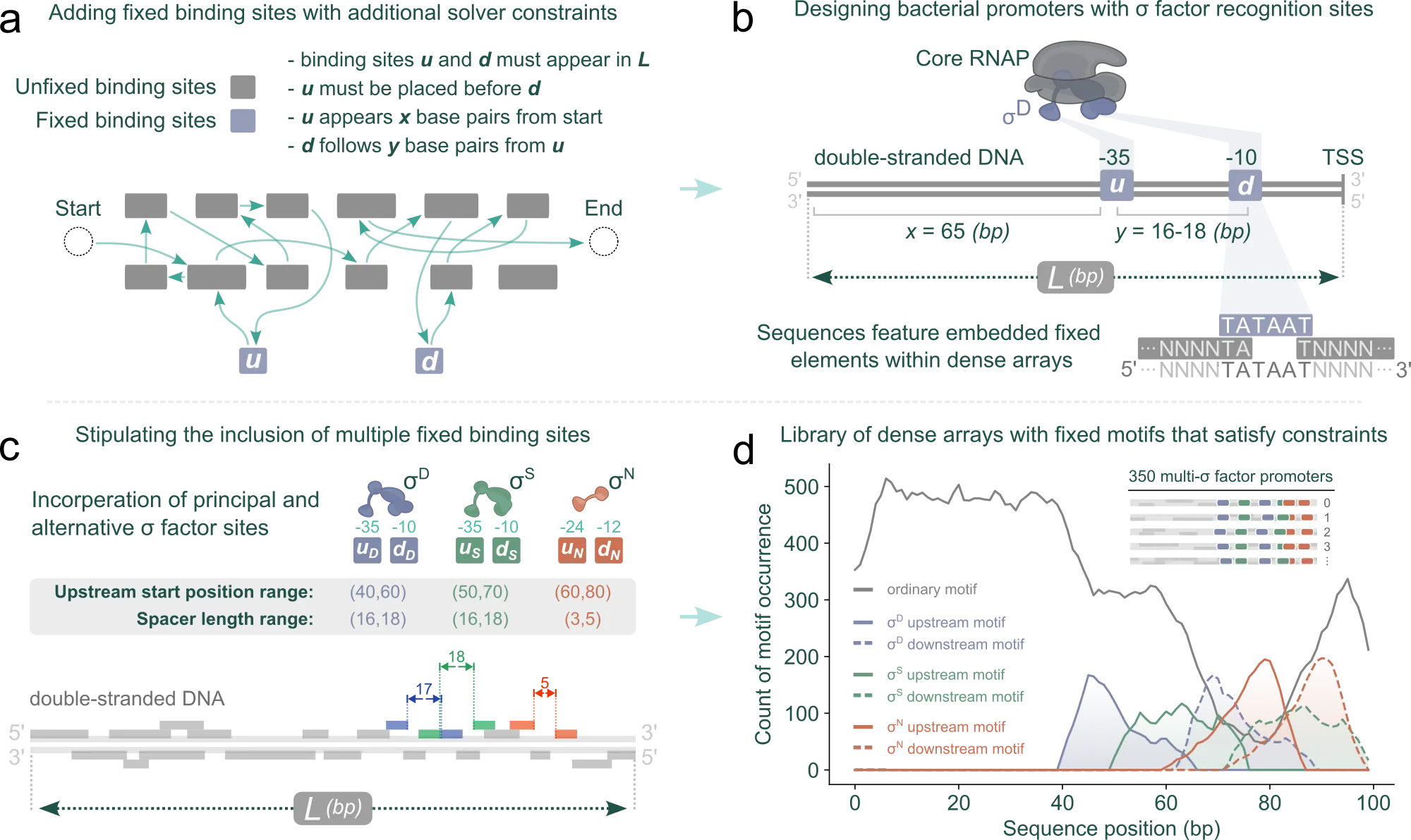
The schematic below explains the generic string-packing principle: many binding sites can be arranged inside a short sequence by treating sequence composition as a constrained graph traversal problem rather than a manual layout exercise.

The DenseGen showcase then shows this principle as an inspectable generation run, with constrained sequence construction and diagnostics that make trajectory and quality tradeoffs visible.
We then extended the same solver with additional constraints to generate bacterial promoter-like sequences, including fixed-position elements and spacing rules, so the packed arrays could be used directly in promoter design workflows.